In silico-prediction of protein–protein interactions network about MAPKs and PP2Cs reveals a novel docking site variants in Brachypodium distachyon | Scientific Reports
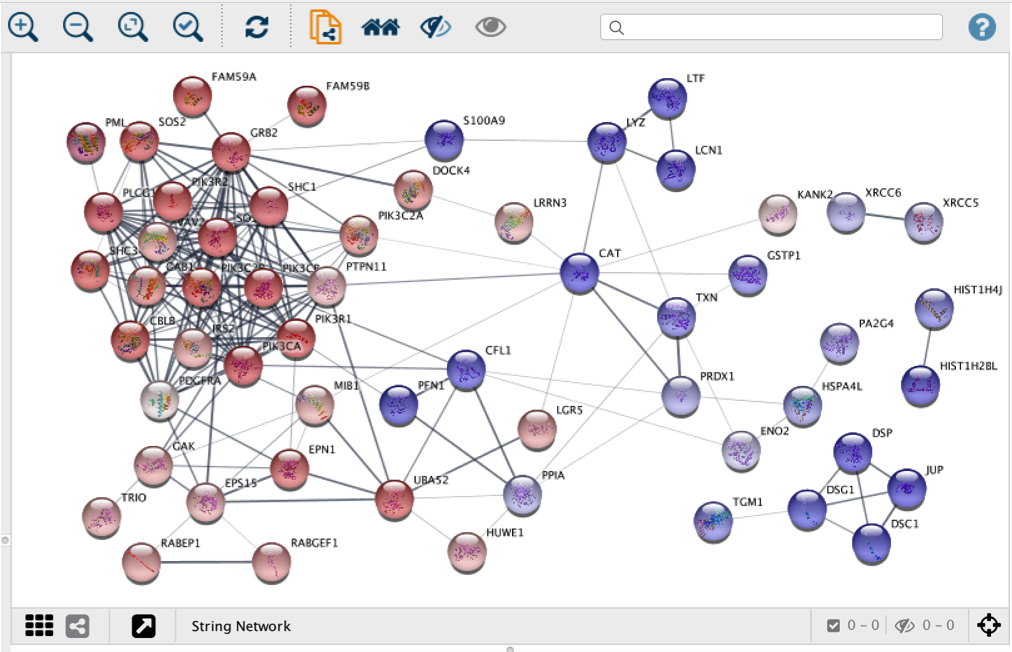
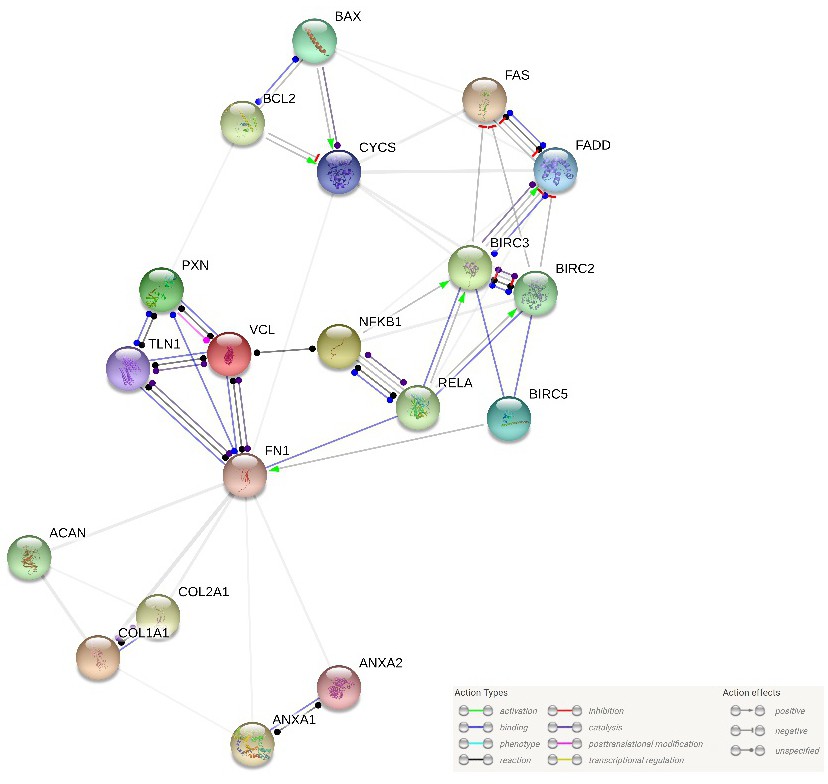
![Plasma proteomic analysis of systemic lupus erythematosus patients using liquid chromatography/tandem mass spectrometry with label-free quantification [PeerJ] Plasma proteomic analysis of systemic lupus erythematosus patients using liquid chromatography/tandem mass spectrometry with label-free quantification [PeerJ]](https://dfzljdn9uc3pi.cloudfront.net/2018/4730/1/fig-5-2x.jpg)
Plasma proteomic analysis of systemic lupus erythematosus patients using liquid chromatography/tandem mass spectrometry with label-free quantification [PeerJ]
![PDF] STRING v11: protein–protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets | Semantic Scholar PDF] STRING v11: protein–protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/1b6d2c08e5b0b7ec50366b0175d266ed37ab7d77/3-Figure1-1.png)
PDF] STRING v11: protein–protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets | Semantic Scholar
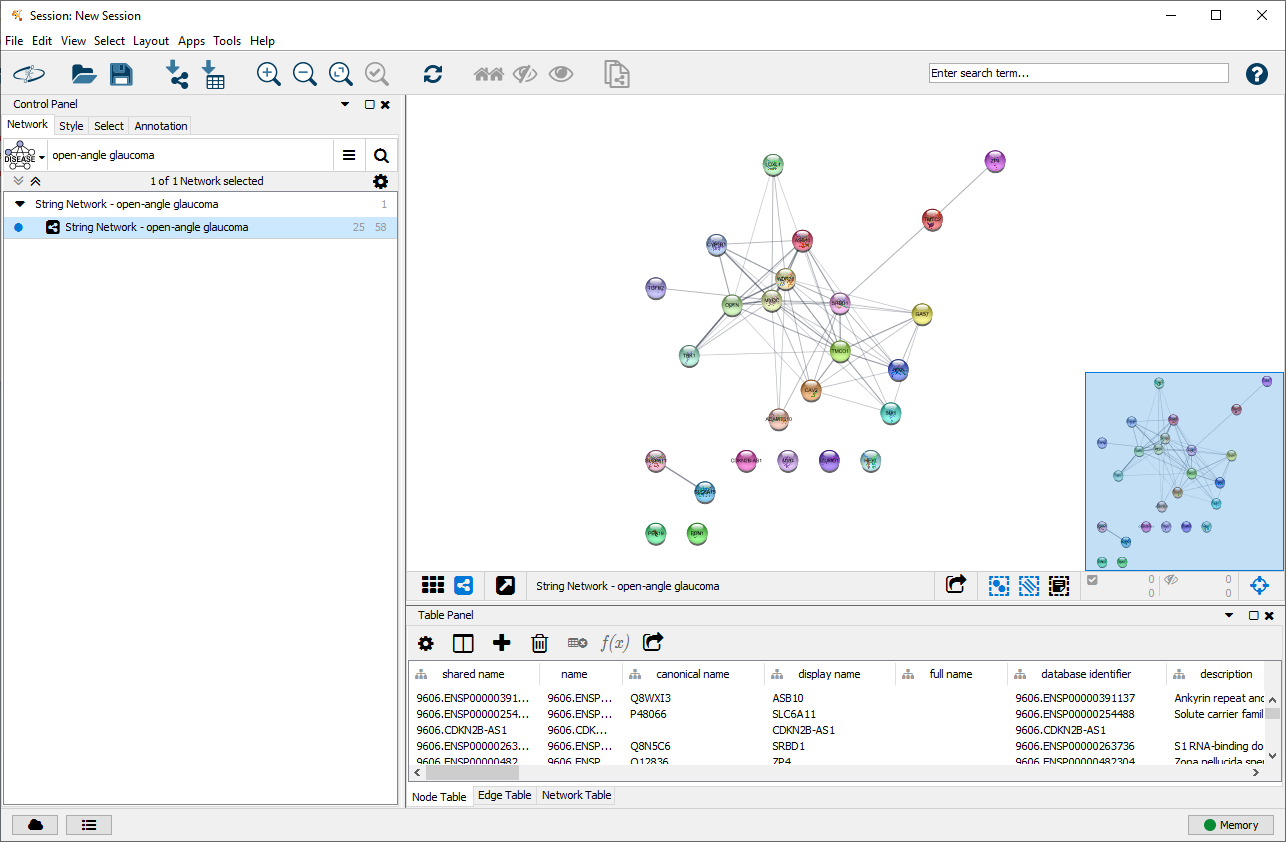
![PDF] The STRING database in 2017: quality-controlled protein–protein association networks, made broadly accessible | Semantic Scholar PDF] The STRING database in 2017: quality-controlled protein–protein association networks, made broadly accessible | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/6e1e6afb314f9c5a24d744252a30aa5efc313571/3-Figure1-1.png)
PDF] The STRING database in 2017: quality-controlled protein–protein association networks, made broadly accessible | Semantic Scholar

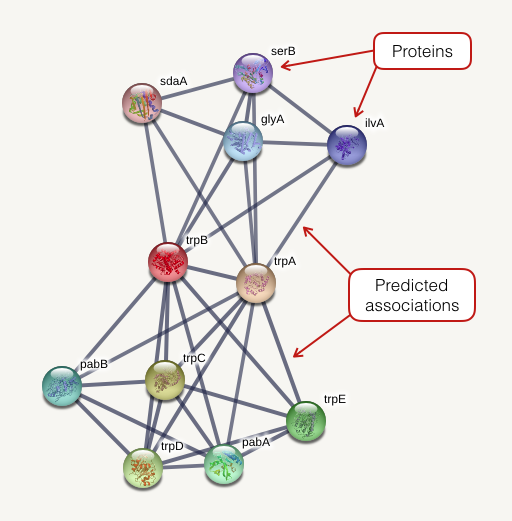
Analysis of protein interaction networks using the STRING system. TAIR... | Download Scientific Diagram

WeiBI (web-based platform): Enriching integrated interaction network with increased coverage and functional proteins from genome-wide experimental OMICS data | Scientific Reports

Insects | Free Full-Text | Evaluation of Total Female and Male Aedes aegypti Proteomes Reveals Significant Predictive Protein–Protein Interactions, Functional Ontologies, and Differentially Abundant Proteins
Identification of Host-Immune Response Protein Candidates in the Sera of Human Oral Squamous Cell Carcinoma Patients | PLOS ONE
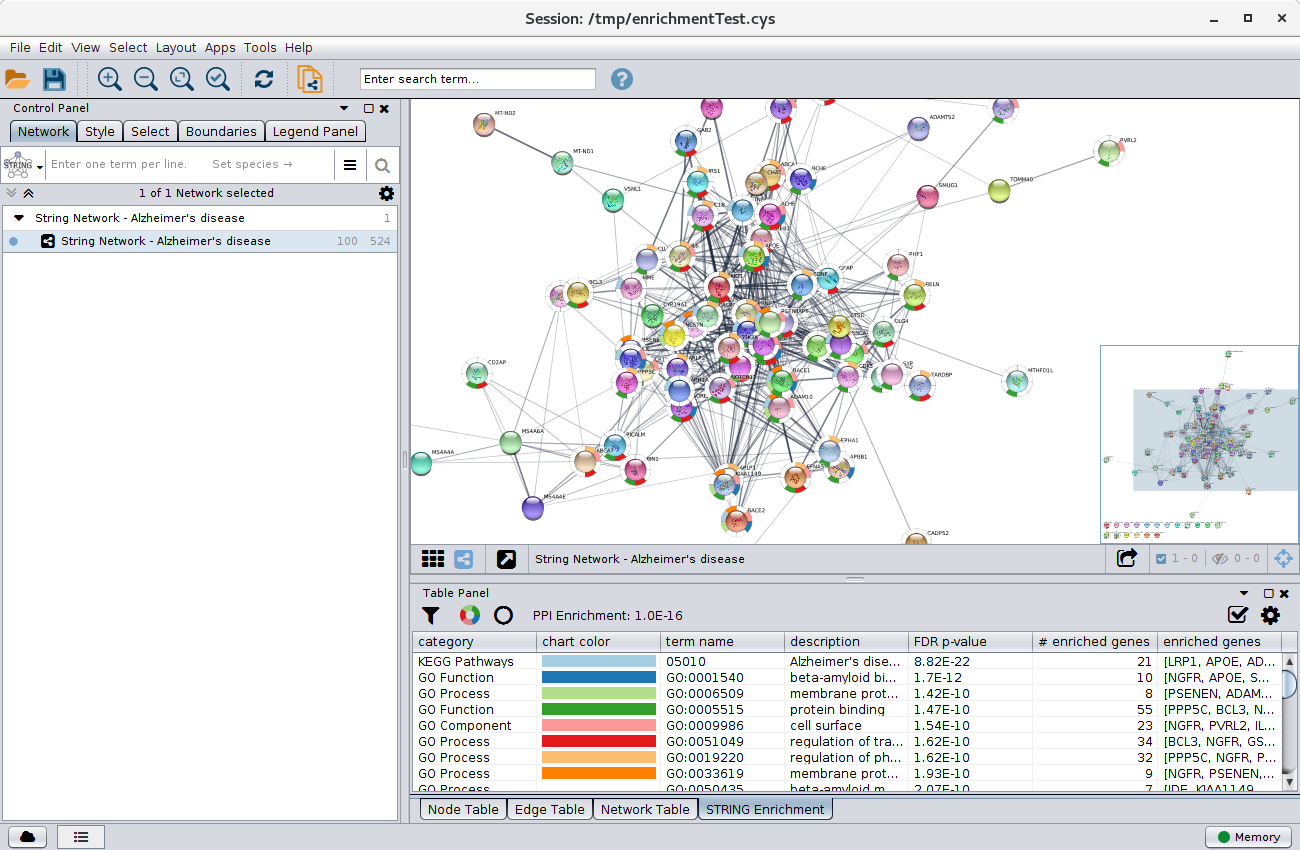
![PDF] The STRING database in 2021: customizable protein–protein networks, and functional characterization of user-uploaded gene/measurement sets | Semantic Scholar PDF] The STRING database in 2021: customizable protein–protein networks, and functional characterization of user-uploaded gene/measurement sets | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/dfd0dd75966b16d12b980f9b964de53f69ee8d08/4-Figure1-1.png)
PDF] The STRING database in 2021: customizable protein–protein networks, and functional characterization of user-uploaded gene/measurement sets | Semantic Scholar
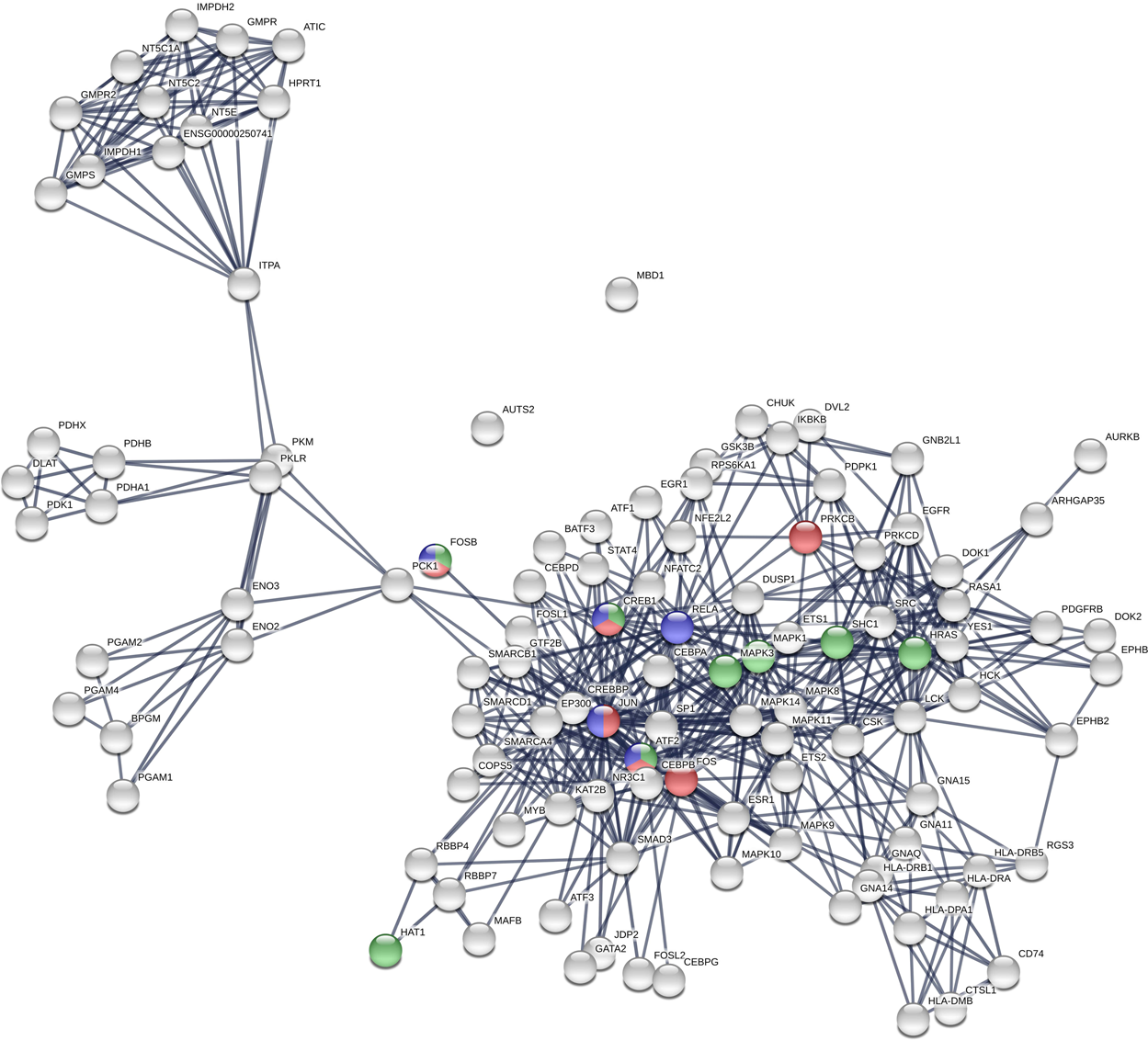
Construction and Analysis of Protein-Protein Interaction Network of Heroin Use Disorder | Scientific Reports