CAGE-Seq Reveals that HIV-1 Latent Infection Does Not Trigger Unique Cellular Responses in a Jurkat T Cell Model | Journal of Virology

Genetic and Epigenetic Features of Promoters with Ubiquitous Chromatin Accessibility Support Ubiquitous Transcription of Cell-essential Genes | bioRxiv

High-resolution annotation of the mouse preimplantation embryo transcriptome using long-read sequencing | Nature Communications
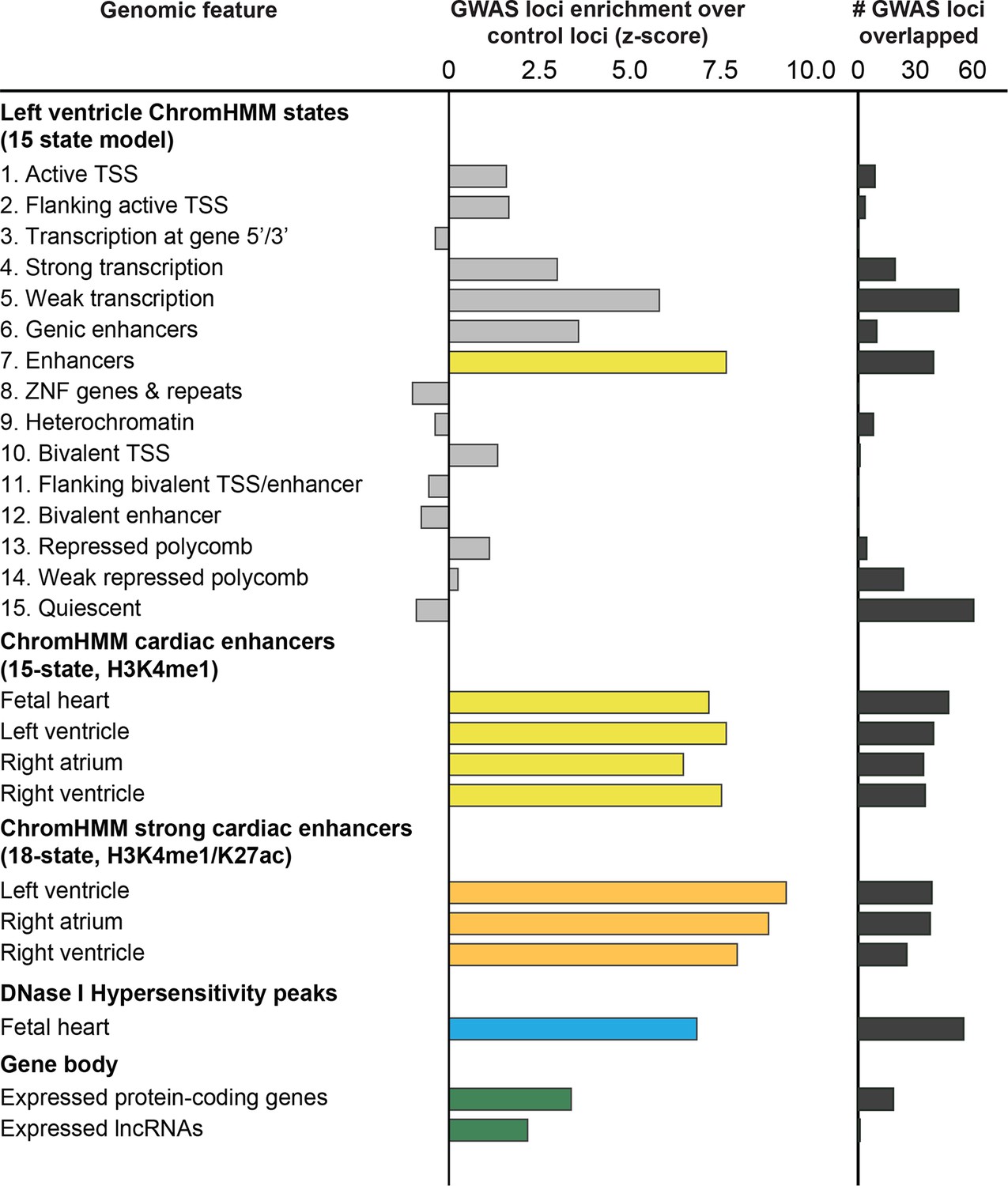
Discovery and validation of sub-threshold genome-wide association study loci using epigenomic signatures | eLife

NET-CAGE characterizes the dynamics and topology of human transcribed cis-regulatory elements | Nature Genetics
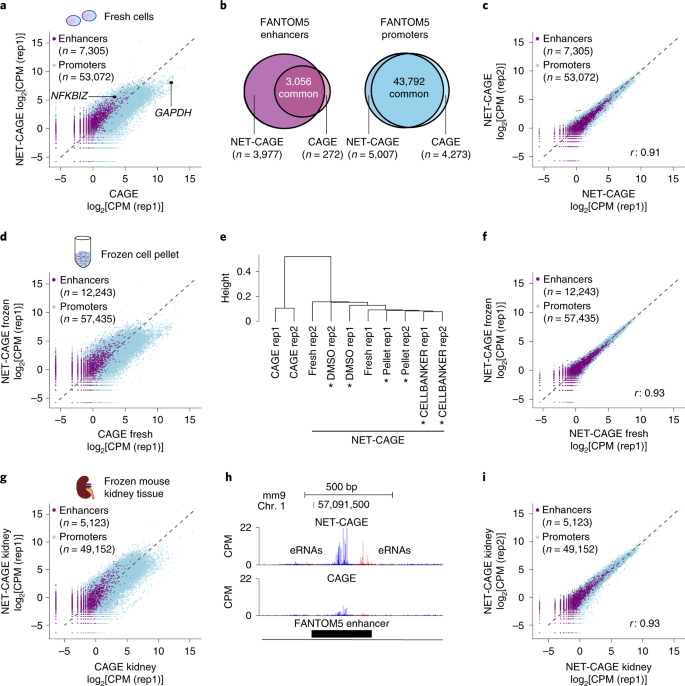
NET-CAGE characterizes the dynamics and topology of human transcribed cis-regulatory elements | Nature Genetics

Cap Analysis of Gene Expression (CAGE): A Quantitative and Genome-Wide Assay of Transcription Start Sites | SpringerLink

NET-CAGE characterizes the dynamics and topology of human transcribed cis-regulatory elements | Nature Genetics

NET-CAGE characterizes the dynamics and topology of human transcribed cis-regulatory elements | Nature Genetics
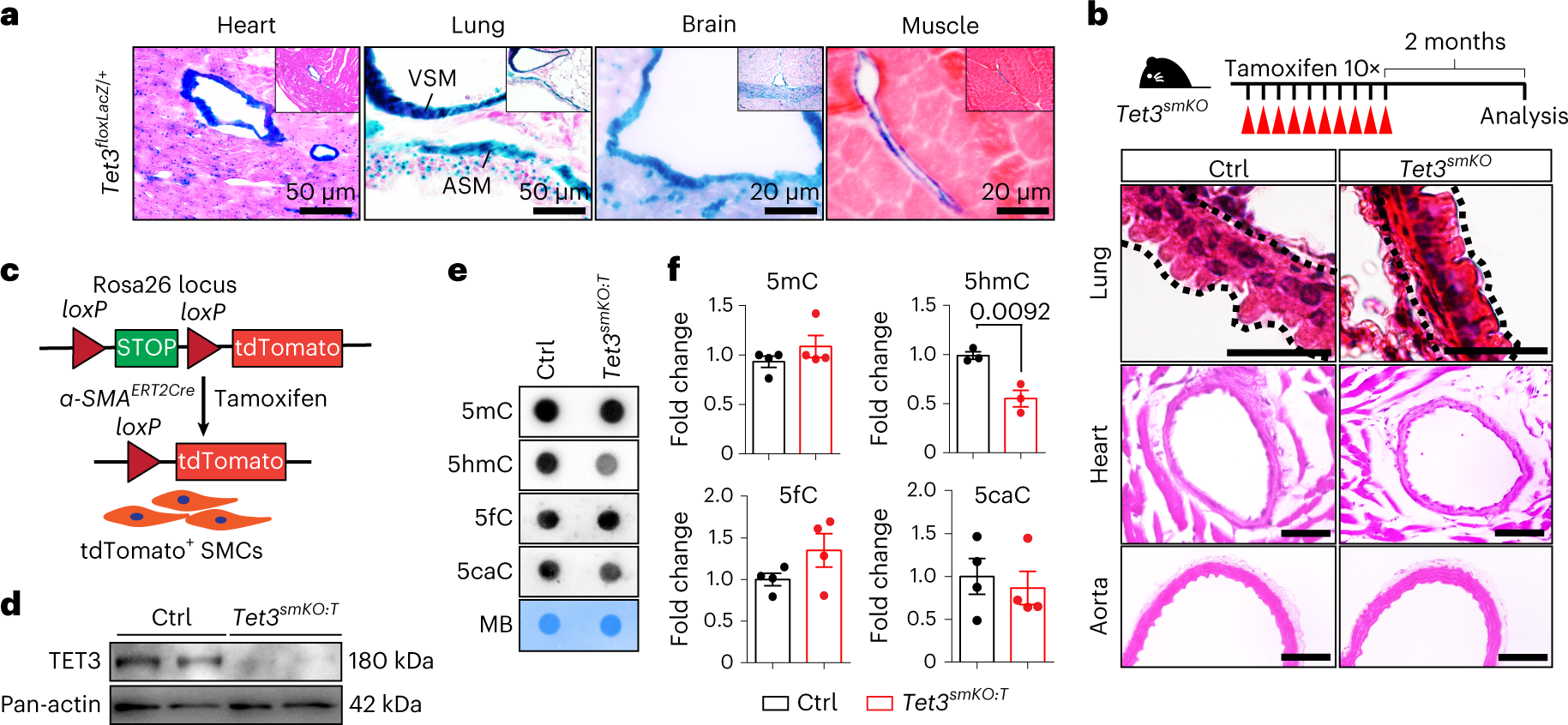
Spurious transcription causing innate immune responses is prevented by 5-hydroxymethylcytosine | Nature Genetics

Discovery and validation of sub-threshold genome-wide association study loci using epigenomic signatures | eLife

Identification of candidate PAX2-regulated genes implicated in human kidney development | Scientific Reports
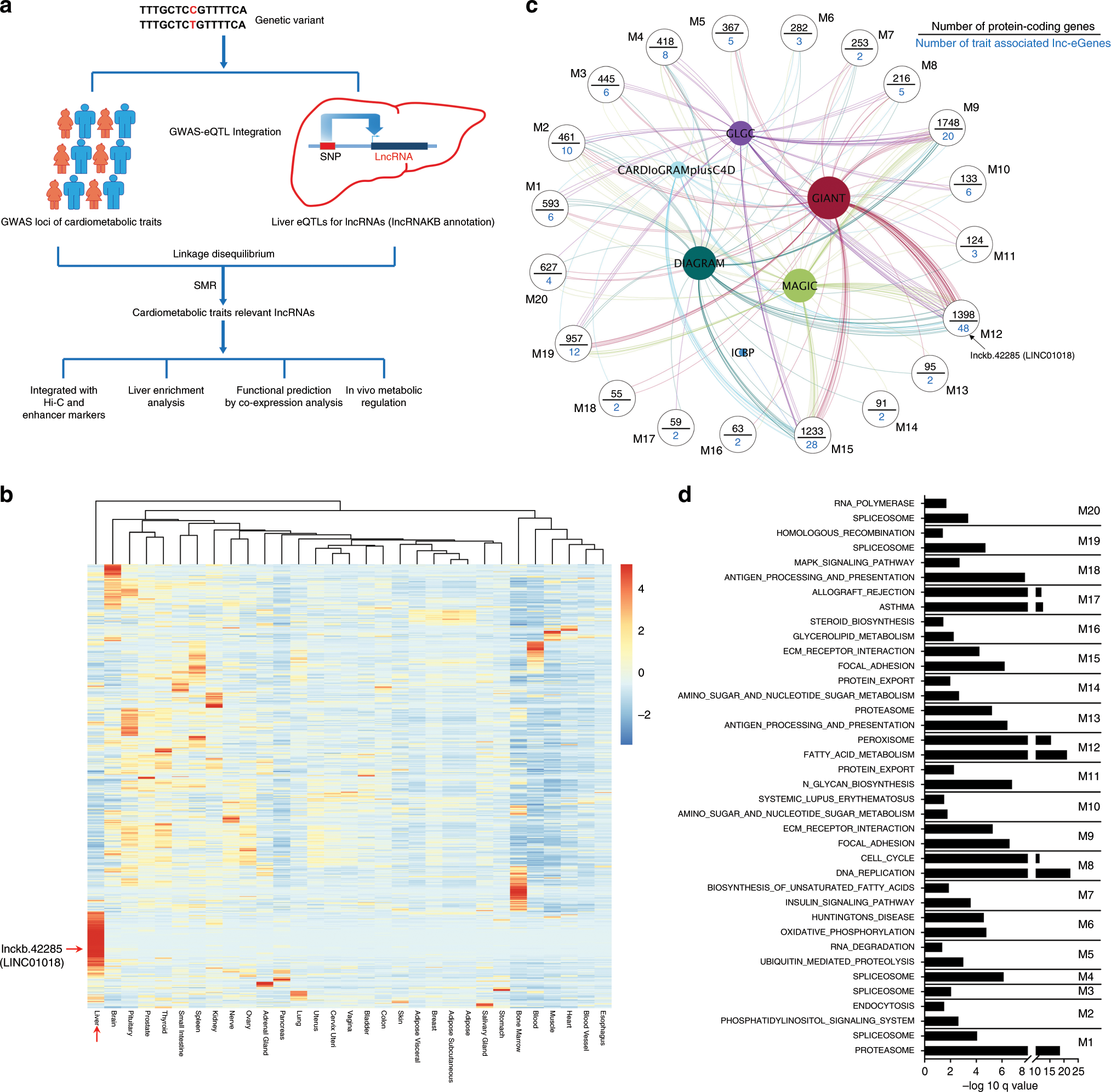
In vivo functional analysis of non-conserved human lncRNAs associated with cardiometabolic traits | Nature Communications